|
3/22/2023 0 Comments Illumina adapter sequences 
This scalable solution has multiple options for sequencing platforms, data analysis, and support to meet virtually all throughput needs. Increase lab efficiency with an easy workflow at as low as 5 hours library preparation workflow and only 1.5 hours of hands-on time. Multiple genes can be analyzed in a single assay, saving time and reducing costs. A multiplexed, polymerase chain reaction (PCR)-based workflow replaces nonspecific hybridization steps, resulting in a high-specificity, high-uniformity amplified library. Library preparation with AmpliSeq for Illumina is fast and simple. Start with either high-quality DNA/RNA samples or use formalin-fixed, paraffin-embedded (FFPE) tissues. Take advantage of low input from as little as 1 ng and flexible input ranges from 1 to 100 ng. 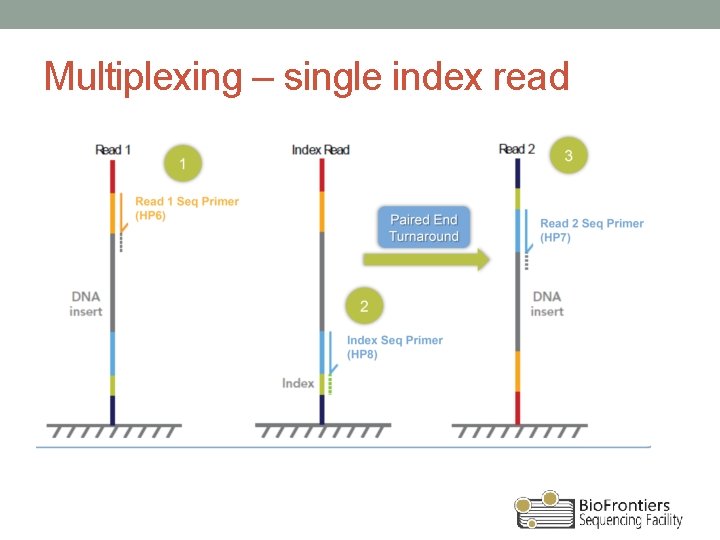
When you run cutadapt you give it the adapter sequence to trim, and this is different for R1 and R2 reads.AmpliSeq for Illumina Library PLUS and Indexes are optimized for use with Illumina sequencers, and generate highly accurate data with an extensive menu of ready-to-use or made to order content. Running cutadapt on small RNA library data The most common application of cutadapt is to remove adapter contamination from small RNA library sequence data, so that's what we'll show here. A high-scoring alignment indicates that the first parts of each read are reverse complements, while the remaining parts of the reads match the respective adapters. The program is not available through TACC's module system but we've installed a copy in our $BI/bin directory. The adapter sequences are prepended to their respective reads, and then the combined read-with-adapter sequences from the pair are aligned against each other. The cutadapt program is an excellent tool for removing adapter contamination. The GSAF website describes the flavaors of Illumina adapter and barcode sequence in more detail Cutadapt trimming 100 bp sequences to 40 or 50 bp), adapter trimming removes differing numbers of 3' bases depending on where the adapter sequence is found. Unlike general fixed-length trimming (e.g. This 3' adapter contamination can cause the "reql" insert sequence not to align because the adapter sequence does not correspond to the bases at the 3' end of the reference genome sequence. The Overrepresented Sequences report, which helps evaluate adapter contamination.ĭata from RNA-seq or other library prep methods that resulted in very short fragments can cause problems with moderately long (50-100bp) reads since the 3' end of sequence can be read through to the 3' adapter at a variable position.But note that different experiment types are expected to have vastly different duplication profiles. The Sequence Duplication Levels report, which helps you evaluate library enrichment / complexity.The Per base sequence quality report, which can help you decide if sequence trimming is needed before alignment.The FastQC reports I find most useful are: Instead, look through the individual reports and evaluate them according to your experiment type. Its "grading scale" (green - good, yellow - warning, red - failed) incorporates assumptions for a particular kind of experiment, and is not applicable to most real-world data. Online documentation for each FastQC reportįirst and foremost, the FastQC "Summary" should generally be ignored.FastQC report for a bad Illumina dataset.FastQC report for a good Illumina dataset.FastQCįastQC is a tool that produces a quality analysis report on FASTQ files. This sequence can also coincide with the binding site for the Index 2 sequencing primer for the optional i5 index (Index 2, purple). The P5 adapter Contains a flow cell binding region (black). The insert sequence (gray) is flanked by two sequencing adapters. This often-overlooked step helps guide the manner in which you process the data, and can prevent many headaches. Figure 1 Structure of a sequencing-ready Illumina-compatible library. The first order of business after receiving sequencing data should be to check your data quality. What is the 2nd sequence in the file /corral-repl/utexas/BioITeam/ngs_course/intro_to_mapping/data/SRR030257_1.fastq? See the Wikipedia FASTQ format page for more information.Įxercise: Examine the 2nd sequence in a FASTQ file For each base, an integer quality score = -10 log(probabilty base is wrong) is calculated, then added to 33 to make a number in the Ascii printable character range. Line 4 is a string of Ascii-encoded base quality scores, one character per base in the sequence.Line 3 is always '+' from GSAF (it can optionally include a sequence description).Line 2 is the sequence reported by the machine.Except for the barcode information, read identifiers will be identical for corresponding entries in the R1 and R2 fastq files. Line 1 is the read identifier, which describes the machine, flowcell, cluster, grid coordinate, end and barcode for the read.GCGTTGGTGGCATAGTGGTGAGCATAGCTGCCTTCCAAGCAGTTATGGGAG 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
